Guanghao(Gary) Wei
MLE @ ByteDance | ex-RA @ Cornell Relax ML Lab

ByteDance Ltd.
1199 Coleman Ave.
San Jose, CA 95110
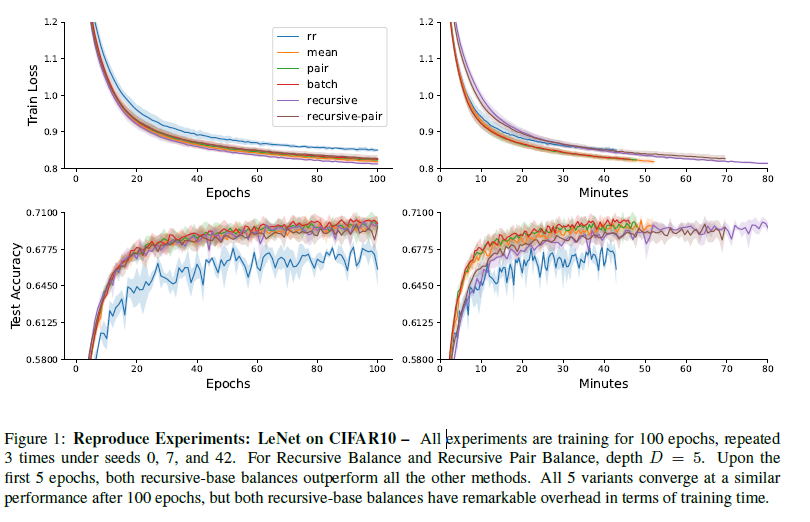
My research interests lie at the intersection of Machine Learning Systems, High Performance Computing, and AI for Science, where I leverage my mathematical and engineering talents to pioneer cutting-edge solutions. I was a Research Assistant at Cornell Relax ML Lab advised by Prof. Chris De Sa on efficient machine learning algorithms and systems. Grounded in mathematical principles, our work aims to expedite large-scale, high-performance machine learning systems that are efficient, parallel, and distributed in real-world settings. Parallel to this, I collaborate on AI-driven molecule generation with talented Ph.D. students.
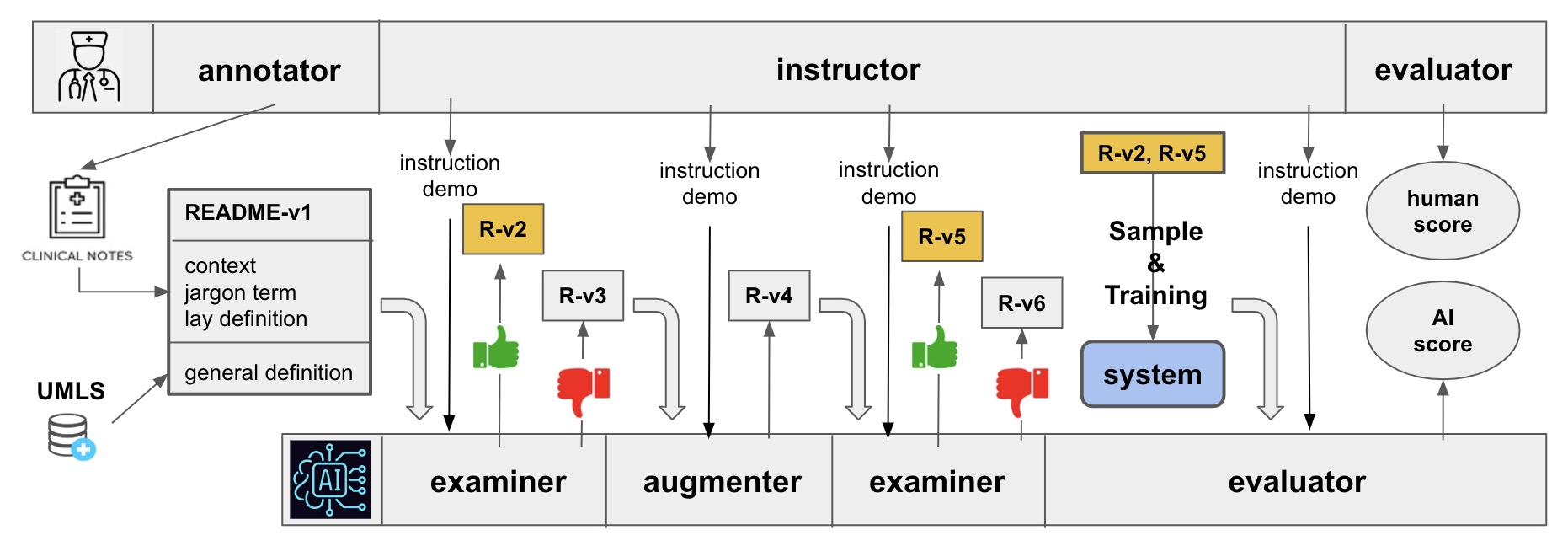
My academic journey has led me to a Master of Engineering in Computer Science at Cornell University. Prior to this, I pursued dual B.S. degrees in Computer Science and Mathematics at the University of Massachusetts Amherst, complemented by a minor in Japanese. My specialization in Mathematics focused on Applied Math and Scientific Computing. Prior to my graduation, I engaged in research at the UMass BioNLP Lab under Prof. Hong Yu, concentrating on bio-medical and clinical NLP applications within Electronic Health Records.
Check out garywei.dev for my personal website!
news
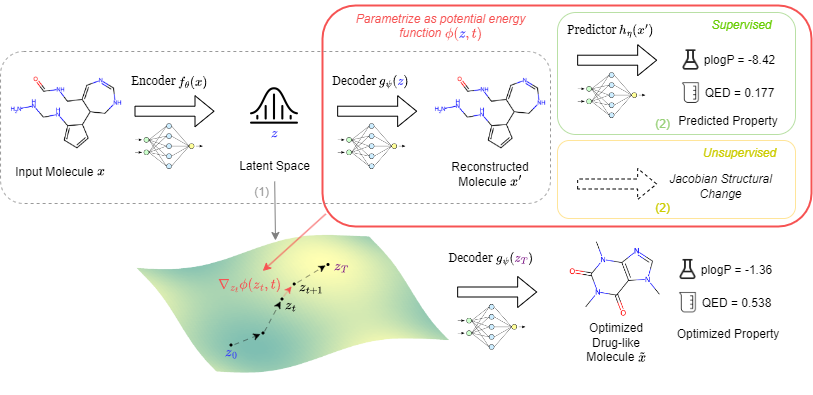
| Sep 26, 2024 | I am excited to share that our paper, “Navigating Chemical Space with Latent Flows,” has been accepted by NeurIPS 2024. |
|---|---|
| Jun 16, 2024 | I am thrilled to announce that our paper, “Navigating Chemical Space with Latent Flows,” received the Spotlight (Top 10%) at the ICML 2024 AI for Science workshop. |
| Feb 29, 2024 | I am happy to continue my position at the Relax ML Lab at Cornell University as a graduate researcher. |
| Dec 31, 2023 | I am delighted to have graduated from Cornell University with a Master of Engineering in Computer Science. |